Serviços Personalizados
Journal
Artigo
Indicadores
-
 Citado por SciELO
Citado por SciELO
Links relacionados
-
 Similares em
SciELO
Similares em
SciELO
Compartilhar
Revista Pan-Amazônica de Saúde
versão impressa ISSN 2176-6215versão On-line ISSN 2176-6223
Rev Pan-Amaz Saude v.1 n.1 Ananindeua mar. 2010
http://dx.doi.org/10.5123/S2176-62232010000100015
ORIGINAL ARTICLE
Phenotypic and genotypic characterization of Serratia marcescens from a Neonatal Unit in Belém, Pará State, Brazil
Raimundo Gladson Corrêa CarvalhoI; Irna Carla do Rosário Souza CarneiroII; Marcelo Sena PinheiroIII; Surama da Costa PinheiroIV; Paulo Sérgio Roffe AzevedoIV; Schirley Dias dos SantosV; Ana Roberta Fusco da CostaVI; Francisco Lúzio de Paula RamosVI; Karla Valéria Batista LimaVI
ICurso de Pós-graduação
em Análises Clínicas, Centro Universitário do Pará,
Belém, Pará, Brasil. Laboratório de Patologia Clínica
e Laboratorial Dr. Paulo Azevedo, Belém, Pará, Brasil
IIDepartamento de Medicina, Universidade Federal do Pará,
Belém, Pará, Brasil
IIICurso de Pós-graduação em Análises
Clínicas, Centro Universitário do Pará, Belém,
Pará, Brasil
IVLaboratório de Patologia Clínica e Laboratorial
Dr. Paulo Azevedo, Belém, Pará, Brasil
VCurso de Pós-graduação em Análises
Clínicas, Centro Universitário do Pará, Belém,
Pará, Brasil. Seção de Bacteriologia e Micologia, Instituto
Evandro Chagas/SVS/MS, Ananindeua, Pará, Brasil
VISeção de Bacteriologia e Micologia, Instituto Evandro
Chagas/SVS/MS, Ananindeua, Pará, Brasil
Endereço para correspondência
Correspondence
Dirección para correspondencia
Original Title: Caracterização fenotípica e genotípica de Serratia marcescens provenientes de Unidade Neonatal de Referência em Belém, Pará. Translated by: American Journal Experts
ABSTRACT
Serratia marcescens has been reported as an important agent of health care-related infections and has been highlighted for presenting a high level of intrinsic resistance to antimicrobials used in neonatology, besides persisting in hospital environments for long periods. In this work, S. marcescens was recovered from colonies in the gastrointestinal tract or late sepsis in newborn infants hospitalized in a Neonatal Unit in Belém. The identification of S. marcescens and the sensitivity test was carried out using a Vitek (BioMérieux) automated system; susceptibility to ertapenem was assessed using e-test strips (Oxoid). Genotyping was executed by ERIC-PCR using the primers ERIC1 (5'-TGAATCCCCAGGAGCTTACAT-3') and ERIC2 (5'-AAGTAAGTGACTGGGGTGAGCG-3'). Twenty-two strains of S. marcescens were recovered: 15 from hemocultures and 7 from surveillance (rectal swab culture). All presented resistance to ampicillin, ampicillin-sulbactam, gentamicin and cephalothin. There were no indications of resistance to ciprofloxacin, imipenem, meropenem or ertapenem. The susceptibility profiles varied for other antibiotics. Eleven amplification patterns by ERIC-PCR were obtained, and two were shared by 14 isolates. It was possible to observe a characteristic polymorphic pattern in the strains from gastrointestinal colonization, except for two cases, which presented genotypic patterns related to cases of sepsis. The data obtained in this work confirm the high level of resistance of S. marcescens against antimicrobials; however, all isolates displayed sensitivity to ciprofloxacin and carbapenemics. Antibiogram and ERIC-PCR typing suggest a dispersion of clones associated with colonization or sepsis among the wards of the Neonatal Unit in the surveyed hospital.
Keywords: Serratia marcescens; Bacterial Typing Techniques; Polymerase Chain Reaction; Bacterial Drug Resistance.
INTRODUCTION
Serratia marcescens, which belongs to the Enterobacteriaceae family, has been reported to be an important agent of health care-related infections (HCRI), specifically standing out due to its dissemination potential and its high level of intrinsic resistance to the drugs used in neonatology, and to antiseptic agents. This pathogen persists for long periods in the hospital environment because it colonizes the skin and the gastrointestinal tract of adults and newborns11,2,6. Therefore, after reporting infections, it is necessary to determine the origin of the pathogen and maintain surveillance, in order to effectively control and/or eradicate the cases.
Many methods have been used for typing epidemic strains of S. marcescens. They involve both phenotypic and genotypic characterizations, and are based on the assumption that related organisms have unique characteristics that distinguish them from non-related organisms. The phenotypic characteristics (e.g., biochemistry, antimicrobial resistance, serotyping, and phage typing) may not be discriminatory, so the use of molecular methods is always necessary for clonal confirmation. The techniques that use pulsed field gel electrophoresis (PFGE) and ribotyping have good discriminatory ability but have the disadvantage of being time-consuming, labor- intensive, and require expensive equipment and reagents9.
Molecular techniques based on the amplification of nucleic acids, such as Enterobacterial Repetitive Intergenic Consensus (ERIC), have been used due to their ease of use, reproducibility, and their agreement with the results obtained by ribotyping5,2,12.
In this study, strains of S. marcescens, isolated during colonization or late sepsis from the Neonatal Unit (NU) of Belém, were evaluated in regards to their antimicrobial resistance profile and genetic characterization.
MATERIALS AND METHODS
DESCRIPTION OF THE INSTITUTION AND SAMPLES
The study was conducted at a NU with a capacity of 103 beds, located at a high complexity hospital in the city of Belém, Pará. The unit consists of an internal nursery (for infants who were born at the hospital), a neonatal intensive care unit (NICU), and an external nursery (EN). The internal nursery is divided in five separate wards: special care (SC), intermediate care (IC), semi-intensive (SI), and transition room (TR). Most newborns referred to the unit are premature with very low weight.
During the study, 675 blood cultures and 75 surveillance cultures were isolated to evaluate the colonization of the intestinal tract by S. marcescens strains.
COLLECTION AND SELECTION OF SAMPLES
Blood cultures were performed on 3 mL of venous blood from each newborn using the automated BACTEC 9120 (Becton Dickinson) system. Surveillance of the colonizing agent within the newborns was performed by cultivating rectal swabs, which were obtained starting on the seventh day of neonate hospitalization and repeated weekly until hospital discharge. The positive primary cultures were subcultured in blood agar (Difco) and MacConckey agar (Difco), and incubated in a bacteriological incubator (Fanen) at 37° C for 24 h. The resulting colonies of fermenting (oxidase negative) Gram-negative bacilli were suspended in 0.45% saline solution, standardized in a colorimeter (BioMerieux) to a concentration equivalent to a 0.5 tube of the Mac-Farland scale (1.5 106 UFC/ mL) and incubated in GN cards for identification by the automated VITEK (BioMerieux) system. The species S. marcescens was selected for this study.
The S. marcescens strain ATCC 8100 and samples not related to the outbreak were used as controls during the genotyping.
SENSITIVITY TEST
The susceptibility profile was evaluated by the automated VITEC (BioMerieux) system, following the manufacturer's recommendations. The susceptibility of S. marcescens to ampicillin (AM), ampicillin-sulbactam (SAM), amikacin (AN), aztreonam (ATM), cefepime (FEP), cefotaxime (CTX), cefoxitin (FOX), ceftazidime (CAZ), cephalothin (KF), ciprofloxacin (CIP), gentamicin (GN), piperacillin-tazobactam (PTZ), sulfa-trimethoprim (STX), imipenem (IPM), and meropenem (MEM) was evaluated. The sensitivity to ertapenem was evaluated with the aid of a disk containing 10 µg of the drug (Oxoide). Inhibition zone readings were performed using a millimeter ruler.
GENOTYPING OF STRAINS BY ERIC-PCR
The bacterial DNA was extracted using the boiling and freezing method (15 min each) and subjected to amplification by polymerase chain reaction (PCR), using the ERIC1 (5'-TGAATCCCCAGGAGCTTACAT-3') and ERIC2 (5'- AAGTAAGTGACTGGGGTGAGCG-3') primers, according to the protocol originally described by Liu et al5. Restriction fragment length polymorphism analysis was conducted after electrophoresis on a 2% agarose gel, followed by visualization on a UV transilluminator.
The development of this study was approved by the Research Ethics Committee of the Instituto Evandro Chagas, protocol CEP/ IEC n° 003/07, on July 14, 2007.
RESULTS
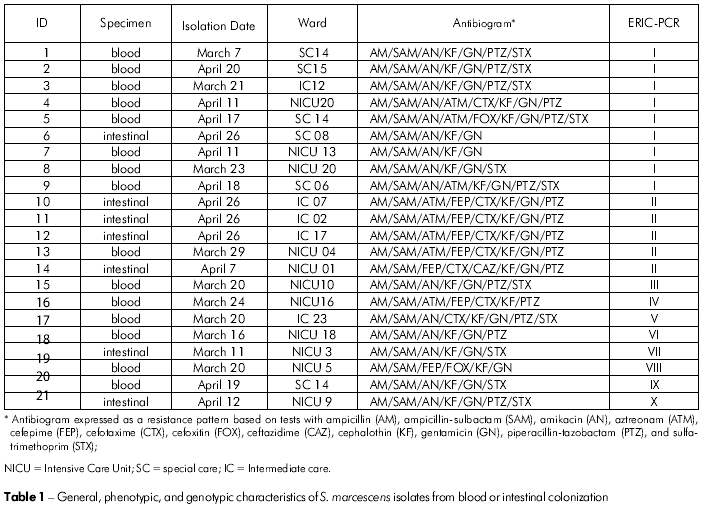
We obtained 22 cultures of S. marcescens, which were recovered from episodes of late neonatal sepsis and colonization. Fifteen of the samples were from neonatal blood cultures, while 7 came from surveillance cultures. All samples showed resistance to AM, SAM, GN, and KF. There was no resistance to CIP, IPM, ertapenem, or MEM. The susceptibility profiles for the other antibiotics tested were variable (Table 1).
We obtained 11 amplification patterns by ERIC-PCR. Eight showed patterns of five or more amplifications between 100 and 3,000 bp (Figure 1), while four samples showed patterns with less than five amplifications (patterns 17, 19, 21). Two polymorphic patterns were shared by 14 isolates (Table 1, Figure 1).
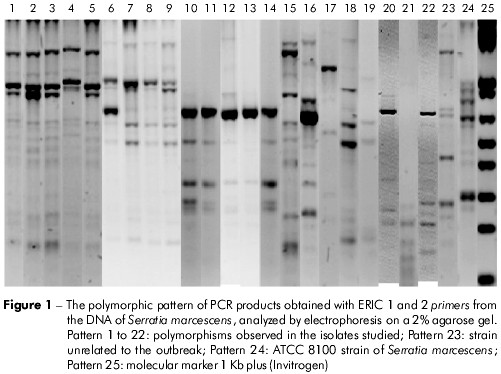
In group I, eight strains were from the blood cultures of newborns hospitalized in different wards of the NU, while the S. marcescens strain designated by the number 6 was recovered from intestinal colonization. In group II, strains 10, 11, 12, and 14 came from rectal swabs (surveillance), while strain 13 was from blood cultures.
DISCUSSION
Serratia marcescens may be associated with sporadic or epidemic infections. It has a high level of resistance to the antibiotics recommended for neonatal therapy11,6 and the antiseptic agents used, which guarantees the survival of this pathogen for long periods in the hospital environment. An important characteristic of this agent is its ability to produce β-lactamase, which gives it resistance to extended-spectrum β-lactam antibiotics and complicates the therapy.
The empirical treatment of HCRI in a NU depends on the time of onset of the infection (early: those seen in the first 48 h of life of the newborn; or late: clinical and laboratory evidence of infection after 48 h of life of the newborn), the previous performance of invasive procedures, the knowledge of local microbes, and the bacterial resistance profile in each hospital1. The protocols may be defined as forms of structured support of technical management, which include definitions of therapeutic goals, temporal sequence of care, and diagnostic and therapeutic strategies.
In this study, all samples of S. marcescens showed resistance to ampicillin and to the associated ampicillin/sulbactam mix. The index of resistance to the aminoglycosides gentamicin and amikacin was also high (Table 1). When evaluating 90 samples of isolated S. marcescens in 1994 at the Hospital of the School of Medicine of Ribeirão Preto, Martinez et al7 observed 99% resistance to ampicillin, 41% resistance to gentamicin, and 21% resistance to amikacin.
In this study, more than 70% (16) of the S. marcescens strains evaluated showed resistance to the β-lactam piperacillin associated with the β-lactamase-inhibitor tazobactam. The combination piperacillin-tazobactam has been a therapeutic alternative for the HCRI caused by Gram-negative rods and β-lactamase-producing anaerobic bacteria present in NICUs, and more recently in NICUs when these microorganisms are isolated during late infections. At the NU evaluated, the combination piperacillin-tazobactam will not be considered for treatment of infections caused by S. marcescens.
The groupings observed after analysis of the polymorphisms obtained by ERIC-PCR for strains obtained from patients hospitalized in different wards configured the spread between rooms of the SC, IC, and NICU. In general, we observed a characteristic polymorphic pattern for the strains originating from gastrointestinal colonization. However, in two cases, strains originating from colonization were also related to cases of sepsis (Table 1). We did not evaluate risk factors associated with septicemia.
Considering that there was no evolution of the infection in newborns with strains 10, 11, 12, and 14, a combination of risk factors is necessary for the development of sepsis, since 80% of them did not present clinical symptoms of septicemia.
Genotyping based on the amplification of repeated elements, similar to ERIC-PCR, has been successfully used for the characterization of various organisms, such as Mycobacterium tuberculosis10, Campylobacters8, and Vibrio cholerae3.
In this study, we evaluated 22 samples of S. marcescens. Seven samples were recovered from the sputum of patients on mechanical ventilators during a pneumonia outbreak, while fifteen were not related to the outbreak5. The data from the ERIC-PCR typing showed two groupings of strains related to the outbreak and distinct profiles among the unrelated strains. These data were confirmed by ribotyping and the profiles of antibiotic resistance.
In this study, differences were observed between the data from ERIC-PCR typing and the pattern of antimicrobial resistance. There was agreement between the typing methods for strains 1, 2 and 3 (Group I); 6 and 7 (Group I); and 10, 11, 12, and 13 (Group II). In other words, 64.29% of the evaluated samples were in agreement. Several factors may justify such a result. First, the susceptibility pattern was not determined following the methods of diskdiffusion, which is standardized and currently recommended by the Clinical and Laboratory Standards Institute (CLSI)4. This technique is important because it is the most fully described and standards have been developed for interpretation of patterns supported by laboratory and clinical data, whose difficulties and improvements are constantly being discussed. Secondly, the profile of bacterial susceptibility may change in a shorter period of time than changes in the patterns of ERIC-PCR. The polymorphic patterns presented in this study retained excellent reproducibility, indicating good discrimination between related and unrelated strains in the outbreak.
In all groupings generated by ERIC-PCR, the sensitivity profiles allowed for subdivision of strains with greater variation of susceptibility to the antibiotics aztreonam, cefotaxime, and cefoxitin. However, this finding is restricted to strains with a high resistance rate.
The S. marcescens strains 8, 19, and 21 showed the same pattern of resistance but differed in the polymorphisms obtained by ERIC-PCR. The low resolution phenotype may originate from the low level of resistance found for these samples, where 100% of the strains showed resistance to the antimicrobials AM, SAM, NA, KF, and GN. Strains 8, 19, and 21 also showed resistance to STX, and was observed in 50% of the evaluated samples. In other words, typing by resistance profile characterization may indicate transmission; however, other methods must be evaluated for confirmation.
We observed the need to increase the number of samples to better evaluate the typing by antibiogram analysis and ERIC-PCR, in addition to developing secondary genotyping of the samples by PFGE, which is currently considered the "golden standard".
In the case of the combination of strains originating from surveillance, S. marcescens acts only as an indicator of transmission. Like S. marcescens, it is possible for other pathogens to be transmitted horizontally and concurrently, which may have been overlooked since it did not represent the focus of the research.
The data obtained in this study confirm the high levels of resistance of S. marcescens strains; however, all evaluated strains showed sensitivity to carbapenems and ciprofloxacin. Typing by ERIC-PCR allowed for grouping of strains, which suggested clones associated with colonization or sepsis spread between the NU wards of the hospital studied.
REFERENCES
1 Agência Nacional de Vigilância Sanitária (BR). Prevenção e controle de infecção hospitalar. Brasília: Ministério da Saúde; 2005.
2 Assadian O, Berger A, Aspöck C, Mustafa S, Kohlhauser C, Hirschl AM. Nosocomial outbreak of Serratia marcescens in a neonatal intensive care unit. Infect Control Hosp Epidemiol. 2002 Aug;23(8):457-6. DOI: 10.1086/502085 [ Links ]
3 Chokesajjawatee N, Zo YG, Colwell RR. Determination of clonality and relatedness of Vibrio cholerae isolates by genomic fingerprinting, using long-range repetitive element sequence-based PCR. Appl Environ Microbiol. 2008 Sep;74(17):5392-401. doi:10.1128/AEM.00151-08 [ Links ]
4 Clinical and Laboratory Standards Institute (USA). Performance standards for antimicrobial disk susceptibility tests; approved standard. 8th ed. Wayne, Pennsylvania; 2003. [ Links ]
5 Liu PYF, Lau YJ, Hu BS, Shir JM, Cheung MH, Shi ZY, et al. Use of PCR to study epidemiology of Serratia marcescens isolates in nosocomial infection. J Clin Microbiol. 1994;32(8):1935-8. [ Links ]
6 Maragakis LL, Winkler A, Tucker MG, Cosgrove SE, Ross T, Lawson E, et al. Outbreak of multidrug-resistant Serratia marcescens infection in a neonatal intensive care unit. Infect Control Hosp Epidemiol. 2008;29(5):418-23. DOI: 10.1086/587969 [ Links ]
7 Martinez R, Gironi RHAR, Santos VR. Sensibilidade bacteriana a antimicrobianos, usados na prática médica - Ribeirão Preto-SP - 1994. Medicina (Ribeirão Preto). 1996 abr-set;29(2-3):278-84. [ Links ]
8 Mouwen DJM, Weijtens MJBM, Capita R, Alonso-Calleja C, Prieto M. Discrimination of enterobacterial repetitive intergenic consensus PCR types of Campylobacter coli and Campylobacter jejuni by Fourier transform infrared spectroscopy. Appl Environ Microbiol. 2005 Aug;71(8):4318-24. DOI:10.1128/AEM.71.8.4318-4324.2005 [ Links ]
9 Parvaz P, Tille D, Meugnier H, Perraud M, Chevallier P, Ritter J, et al. A rapid and easy PCR-RFLP method for genotyping Serratia marcescens strains isolated in different hospital outbreaks and patient environments in the Lyon area, France. J Hosp Infect. 2002 Jun;51(2):96-105. [ Links ]
10 Sechi LA, Zanetti S, Dupré I, Delogu G, Fadda G. Enterobacterial repetitive intergenic consensus sequences as molecular targets for typing of Mycobacterium tuberculosis strains. J Clin Microbiol. 1998 Jan;36(1):128-32. [ Links ]
11 Villari P, Crispino M, Salvadori A, Scarcella A. Molecular epidemiology of an outbreak of Serratia marcescens in a neonatal intensive care unit. Infect Control Hosp Epidemiol. 2001 Oct;22(10):630-4. DOI: 10.1086/501834 [ Links ]
12 Vries JJ, Baas WH, Van der Ploeg K, Heesink A, Degener JE, Arends JP. Outbreak of Serratia marcescens colonization and infection traced to a healthcare worker with long-term carriage on the hands. Infect Control Hosp Epidemiol. 2006 Nov;27(11):1153-8. DOI: 10.1086/508818 [ Links ]
 Correspondência
/ Correspondence / Correspondencia:
Correspondência
/ Correspondence / Correspondencia:
Karla Valéria Batista Lima
Instituto Evandro Chagas,
Seção de Bacteriologia e Micologia
Rodovida BR 316, km 7, s/no,
Levilândia
CEP: 67030-000
Ananindeua-Pará-Brasil
Recebido em / Received / Recibido en: 30/07/2009
Aceito em / Accepted / Aceito en: 01/10/2009










 texto em
texto em