ORIGINAL ARTICLE | ARTIGO ORIGINAL | ARTÍCULO ORIGINAL
Geographic dispersion of Phyllostomidae family (Chiroptera) based on Cytochrome b sequences
Dispersão geográfica da família Phyllostomidae (Chiroptera) baseado nas sequências do cifocromo b
Dispersión geográfica de la familia Phyllostomidae (Chiroptera) basada en las secuencias del cifocromo b
]]> Gustavo Moraes HolandaI; Edivaldo Herculano Correa de OliveiraII; Nelson Antonio Bailão RibeiroIII
ISeção de Arbovirologia e Febres Hemorrágicas, Instituto Evandro Chagas/SVS/MS, Ananindeua, Pará, Brasil
IISeção de Meio Ambiente, Instituto Evandro Chagas/SVS/MS, Ananindeua, Pará, Brasil. Universidade Federal do Pará, Belém, Pará, Brasil
IIICentro de Inovações Tecnológicas, Instituto Evandro Chagas/SVS/MS, Ananindeua, Pará, Brasil. Laboratório de Biologia Molecular, Centro de Ciências Biológicas e da Saúde, Universidade Estadual do Pará, Belém, Pará, Brasil
Correspondence
Endereço para correspondência
Dirección para correspondencia
ABSTRACT
]]> The Chiroptera order is one of the most successful species of mammals with a wide geographical distribution. This order has been traditionally divided into two suborders, Microchiroptera and Megachiroptera, and the family Phyllostomidae is included in the suborder Microchiroptera. However, studies with molecular analysis show a different classification in two different suborders: Yangochiroptera and Yinpterochiroptera. Studies with various species describe a wide dispersal of these animals from Central America to South America and specimens of different places, creating new karyotypes and different nucleotide sequences, especially in the widely known Cytochrome b gene. In this study, we analyzed a phylogeographic dispersion of the Pyllostomidae family using the mitochondrial Cytochrome b gene, a possible dispersion pattern for family and new evolutionary proposals. All the sequences were obtained from the online database (GenBank) and the analysis and formation of phylogenetic trees were performed by maximum parsimony and maximum likelihood methods. Some dispersion patterns were observed for species of genus Carollia and Glossophaga in individual analysis and other species pattern of dispersion from South to West. But in general analysis, a pattern of dispersal to the North of the American Continent was evidenced for the family, following South America to Central America, despite many landforms that could cause speciation of some genera such as isolation by the Andes mountains. Further analysis, with a greater number of specimens from different locations, must be done to confirm this theory.Keywords: Phylogeography; Chiroptera; Genes; Cytochromes b.
RESUMO
A ordem Chiroptera é uma das espécies de mamíferos mais bem sucedidas com uma grande distribuição geográfica. Essa ordem foi tradicionalmente dividida em duas subordens, Microchiroptera e Megachiroptera, e a família Phyllostomidae está incluída na primeira. No entanto, estudos com análise molecular mostram uma classificação diferente em duas subordens distintas: Yangochiroptera e Yinpterochiroptera. Os estudos com várias espécies descrevem uma grande dispersão desses animais da América Central para a América do Sul e espécimes de vários lugares, a criação de novos cariótipos e sequências de nucleotídeos diferentes, especialmente no gene citocromo b amplamente conhecido. Neste estudo, analisou-se uma dispersão filogeográfica da família Pyllostomidae usando o gene mitocondrial citocromo b, um possível padrão de dispersão para essa família e novas propostas evolutivas. Todas as sequências foram obtidas a partir da base de dados on-line (GenBank), a análise e a formação de árvores filogenéticas foram realizadas pelos métodos de máxima parcimônia e de máxima verossimilhança. Foram observados alguns padrões de dispersão de espécies do gênero Carollia e Glossophaga na análise individual e outro padrão de dispersão de espécies do sul ao oeste. Porém, na análise geral, um padrão de dispersão para o norte do Continente Americano foi evidenciado para a família, depois da América do Sul à América Central, apesar de muitos acidentes geográficos causarem especiação de alguns gêneros, tais como o isolamento das montanhas dos Andes. Uma análise mais aprofundada, com um maior número de amostras de diferentes locais, deve ser feita para confirmar esta teoria.
Palavras-chave: Filogeografia; Quirópteros; Genes; Citocromos b.
RESUMEN
El orden Chiroptera es una de las especies de mamíferos de mejores resultados y con una gran distribución geográfica. Ese orden ha sido tradicionalmente dividido en dos subórdenes, Microchiroptera y Megachiroptera, y la familia Phyllostomidae está incluida en la primera. Sin embargo, estudios con análisis molecular muestran una clasificación diferente en dos subórdenes distintas: Yangochiroptera e Yinpterochiroptera. Los estudios con varias especies describen una gran dispersión de esos animales de América Central para América del Sur y especímenes de varios lugares, la creación de nuevos cariotipos y secuencias de nucleótidos distintos, especialmente en el gen citocromo b ampliamente conocido. En este estudio se analiza una dispersión filogeográfica de la familia Pyllostomidae usando el genoma mitocondrial citocromo b, un posible estándar de dispersión para esa familia y nuevas propuestas evolutivas. Todas las secuencias se obtuvieron a partir de la base de datos online (GenBank), el análisis y la formación de los árboles filogenéticos se realizaron por los métodos de máxima parsimonia y máxima verosimilitud. Se observaron algunos estándares de dispersión de especies del género Carollia y Glossophaga en el análisis individual y otro estándar de dispersión de especies del sur al oeste. Sin embargo, se evidenció en el análisis general, un estándar de dispersión hacia el norte del Continente Americano para la familia, después de América del Sur a América Central, a pesar de que muchos accidentes geográficos causan la especiación de algunos géneros, como el aislamiento de las montañas de los Andes. Un análisis más profundo, con un número de muestras de diferentes locales más grande debe hacerse para confirmar esta teoría.
Palabras clave: Filogeografía; Quirópteros; Genes; Citocromos b.
]]> INTRODUCTION
Bats, classified in the order Chiroptera (cheir = hand; pteron = wing), are the only mammals that exhibit morphological and physiological adaptations for true powered flight. Their membranous wings extend between fingers, constituting dactilopatagium, which allow them flap the wings and fly. Other features that distinguish bats are the presence of membranes in the lower limbs called uropatagium and above the lower limbs, called protopatagium1,2.
The order has a wide geographical distribution, found on all continents, except in the Polar Regions. Among all continents, the South and Central America have the high diversity of species, in contrast to North America and northern Eurasia, which have a low diversity, due to increasing latitude and consequently reducing the temperature3,4,5,6,7,8.
Over the decades, the classification of Chiroptera changed several times. The traditional classification recognized two suborders: Megachiroptera and Microchiroptera. Simmons9 has described a total of 18 families, 202 genera and 1,120 species.
The Megachiroptera is composed of a single family Pteropodidae, which has 190 species divide into four subfamilies: Pteropodinae, Harpyonycterinae, Nyctimeninae and Macroglossinae. Microchiroptera has 930 species divide into 17 families: Rhinolophidae, Hipposideridae, Megadermatidae, Rhinopomatidae, Phyllostomidae, Craseonycterinae, Emballonuridae, Nycteridae, Myzopodidae, Mystacinidae, Molossidae, Vespertilionidae, Mormoopidae, Noctilionidae, Furipteridae, Thyropteridae and Natalidae9.
In contrast to the traditional morphological classification, recent cytogenetic and molecular studies have proposed a different classification for bats. Some authors have divided the order into two new suborders Yangochiroptera (consisting of Microchiroptera, except Rhinopomatidae, Rhinolophidae and Megadermatidae) and Yinpterochiroptera (consisting of Megachiroptera and families Rhinopomatidae, Rhinolophidae and Megadermatidae)10,11,12.
Among the several bat families that have been found in Brazil, the most common is the Phyllostomidae (from greek: phyllo = leaf + stoma = mouth), which is composed by a variety of bats that have different eating habits, such as insectivorous, frugivorous, carnivorous, herbivorous, nectarivorous, omnivorous and hematophagous2,13.
The Phyllostomidae family is divided into seven subfamilies: Desmodontinae, Glossophaginae, Phyllostominae, Carolliinae, Stenodermatinae, Phyllonycterinae and Brachyphyllinae and more than 160 species are grouped in 48 genera9. The family was classified in Microchiroptera or Yangochiroptera10,11,12.
The main morphological feature is an appendix in the format of a shaped spearhead, also called "leaf nose", except for Desmodontinae subfamily, and the species Cenfurio senex (the structure is reduced or absent). Its species are mostly adapted to low altitudes in the tropical and subtropical regions of the New World9. They can be found from the Southeastern United States to northern of Argentina13.
At the beginning of this century, there was a large number of taxonomic and systematic revisions in several groups of bats present in Brazil, proposing new classifications between families and genera of Chiroptera9,14,15,16,17.
]]> According to Baker18 there are 49 genera in three subfamilies, Koopman19 recognized 49 genera in eight subfamilies, McKenna and Bell20 recognized 48 genera in four subfamilies, Nowak13 recognized 52 genera in five subfamilies and Wetterer et al16 recognized 53 genera in seven subfamilies. Again Baker et al21 proposed another classification with 11 subfamilies, 57 genera, ten tribes and seven unrated taxa.Since the 1970s, several molecular techniques, such as electrophoresis, polymerase chain reaction (PCR), DNA sequencing, proteomics and others were used in several studies, mainly in phylogenetic14,21,22. These techniques can be used to analyze nuclear DNA and DNA from organelles, allowing the measurement of phylogeographic parameters, gene flow and it's inter and intraspecific historical relationships14,17,21.
Currently, one of the most widely used markers for studying phylogeny and phylogeography is the mitochondrial DNA (mtDNA), which is circular, haploid and matrilineal (maternal inheritance pattern). It is found in the mitochondrial lumen with thousands of copies per cell, almost all of the genome is involved in coding functions and the existence of pseudogenes, introns and repetitive DNA is rare23,24. One of the suitable features for the genetic analysis is the ability to accumulate base substitutions, insertions and deletions at a rate of five to ten times faster than nuclear DNA, due to low replication fidelity, no repair mechanisms, corrosive oxygen-rich environment and no DNA compaction23. The phylogeny based on mtDNA allows a good description of the geographical distribution, genetic distances and divergence times between lineages1.
The most widely used marker is the Cytochrome b gene that has one of the best resolutions for phylogenetic studies in mammals with divergence times from four to 44 million years1,25. The gene contains codons with fast and slow evolution, conserved regions and variable regions26. It has been widely used in analyzes that involves geographic distribution of genetic lineages, evolution, reproductive isolation, origin, distribution, biodiversity maintenance and the emergence of new species27. Based on its variability, Cytochrome b gene is considered appropriate to elucidate relationships to a genus level, the saturation may complicate at higher taxonomic levels14.
Phylogeography studies using molecular data (usually mtDNA and microsatellite) can propose a geographic evolution (based on vicariance, diversification of species and patterns of migration) and demographic history (fixation of species, population growth and decline)3.
Among several works about the phylogeography of the Phyllostomidae family, little is known of its evolution, dispersal and establishment of the species, in general perspective. Most studies use only a few genera, species or small groups, never with a general group, using various genera3,4,5,6,7,8.
In Phyllostomidae, only Leptonycteris curasoae, Leptonycteris nivalis, Choernycteris mexicana and Platalina genovensium needs a migratory behavior. In other families, few species have this behavior and most are classified in the Vespertillionidae family28. This migration, in some species, may be greater in females than in males and changes may occur especially in Neotropics wet and dry seasons29. What differentiates bats from birds, in the migratory aspect, is that bats hibernate in caves and mines during the winter and birds migrate to different locations to get out of the food shortages and climate change30,31.
Bats from North America and South America have been exchanging genetic information and fauna over a millions years, compounding the tropical fauna of mammals in the Americas5. The dispersion has occurred in regions that had a higher amount of food, especially in autumn season32.
Thus, the aim of this study was to analyze the evolutionary and phylogeographic dispersal involving species of bats of the Phyllostomidae using the mitochondrial gene, Cytochrome b, and suggesting a possible dispersion pattern for the family.
]]> MATERIALS AND METHODS
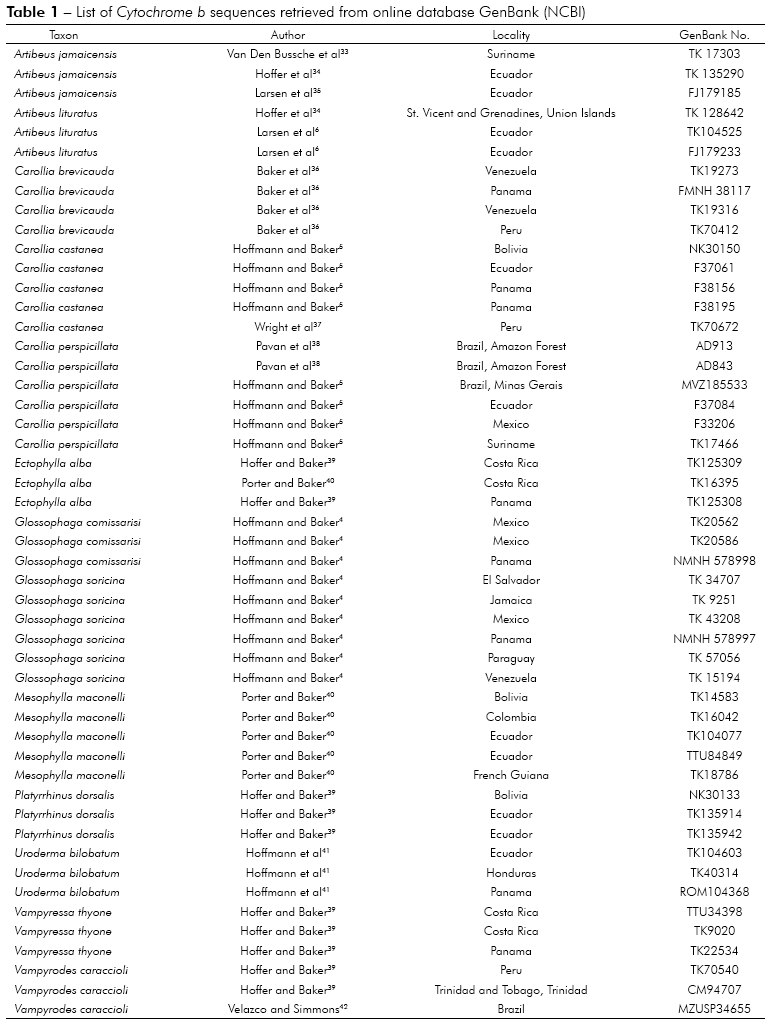
All procedures were performed at Instituto Evandro Chagas in the Centro de Inovações Tecnológicas. The DNA sequences of Cytochrome b gene were obtained directly from GenBank (NCBI) (http://www.ncbi.nlm.nih.gov/genbank). It was retrieved 50 complete Cytochrome b sequences of 1,140 bp for a variety of localities throughout the American Continent (Table 1). It was only used sequences with the locality referred in the original articles.

The numbers of retrieved sequences were low because many sequences were not in the referenced article, others have not been published in articles and others did not mention the sites that the specimens were captured.
PHYLOGENETIC ANALYZES
In order to observe genetic variations, the nucleotide sequences were aligned and compared using BioEdit 7.0 software. For phylogenetic tree formation, it was used the maximum parsimony and maximum likelihood methods. Analysis were performed considering all nucleotides and codons substitutions with heuristic search in PAUP 4.0b10 software using the tree bisection-reconnection (TBR). The bootstrap value was expected in one thousand replications using the heuristic test. In the maximum likelihood analysis were used GTR nucleotide substitution models and TBR with gamma frequencies chosen by software J Model Test 2.1.3. Both analysis were based on the parameters described in Baker21.
It was used the genus Mormoops from the Mormoopidae as outgroup because it is considered one of the closest to the Phyllostomidae according to Baker et al21 and Jones et al17. Intraspecific analysis were performed for each species and interspecific with all genera retrieved from GenBank (Table 1).
]]>RESULTS
In the maximum parsimony analysis, from 1,140 characters, 651 were invariant, 455 were informative and 34 uninformative.
It could be observed seven distinct groups without evolutionary information between them, but internally for the genus Artibeus, it was observed that Artibeus jamaicensis had basal status in relation to Artibeus lituratus. The specimen of Carollia brevicauda had basal status in relation to specimens of the same species and to Carollia pespecillata. Regarding the species Ectophylla alba, the specimen that was obtained from Panama had a basal status compared the ones in Costa Rica. The species Glossophaga commissarisi, from Panama also has basal status relative to all other species from Glossophaga genus (Figure 1).
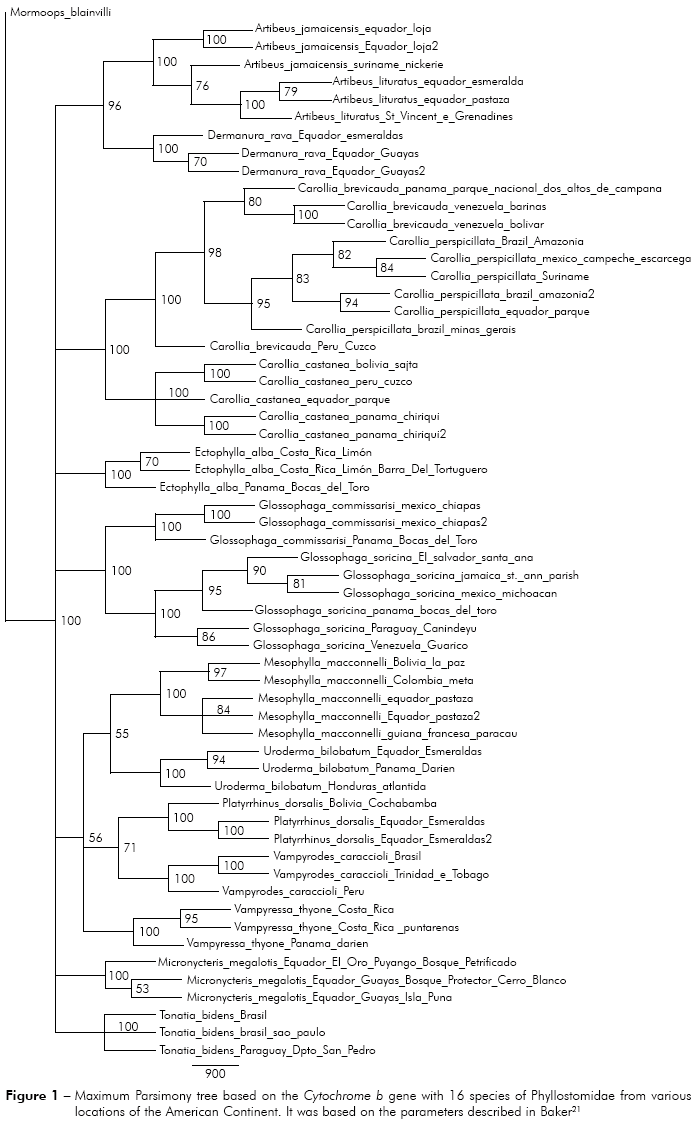
In groups 5, 6, 7 and 8 the specimens with basal status are: Uroderma bilobatum from Honduras, Vampyrodes caraccioli from Peru, Vampyressa thyone from Panama and Micronycteris megalotis from Ecuador (specifically in El Oro, Puyango, Petrified Forest), respectively (Figure 2).
]]>
Regarding the analysis by maximum likelihood, three groups were formed, one with basal status in the family, having the specimens of Micronycteris megalotis, following two distinct groups composed of group 1 (with Tonatia, Glossophaga and Carollia) and group 2 by the remaining genus, with the most basal status for Ectophylla alba.
Artibeus jamaicensis - It was analyzed specimens from two different localities (Suriname and Ecuador). In maximum parsimony and maximum likelihood, the species was dispersed from East to West of South America, in which the specimens from Ecuador are more derived in the phylogenetic analysis than from Suriname.
Artibeus lituratus - It was analyzed specimens from two different localities (St. Vincent and the Grenadines and Ecuador). In maximum parsimony and maximum likelihood, the species were dispersed from East to West and to the South of South America, and the specimens of Ecuador are more derived than those of Saint Vincent and the Grenadines.
Carollia brevicauda - Specimens from three different locations were analyzed (Peru, Panama and Venezuela). In maximum parsimony and maximum likelihood, the species were dispersed from South to North of America, and the specimens of Venezuela are more derived than those from Panama and these are more derived than those from Peru.
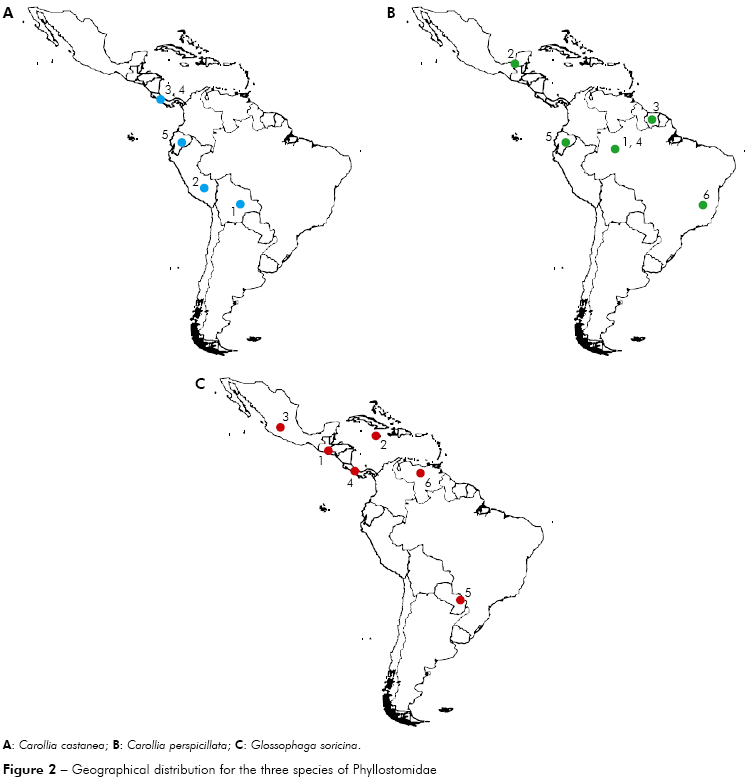
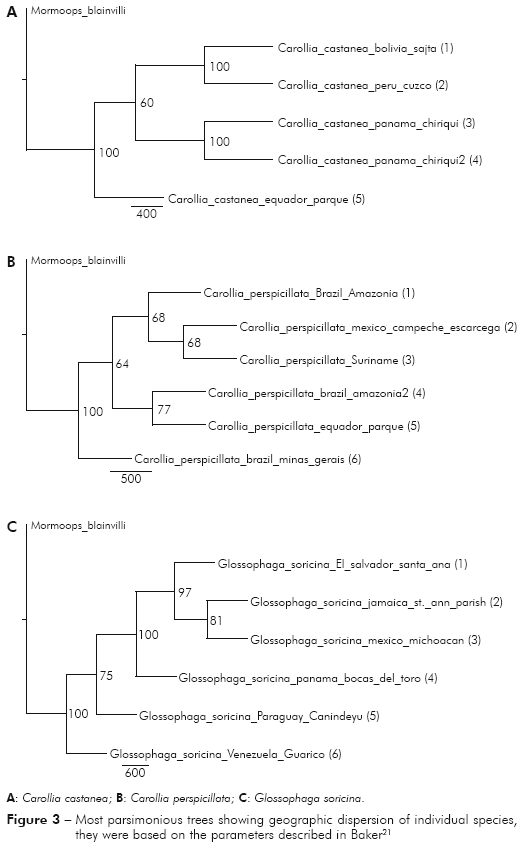
Carollia castanea - Four different locations were analyzed (Peru, Ecuador, Bolivia and Panama). In maximum parsimony the specimens located in Ecuador are more basal, followed by two derived groups, one consisting of specimens from Bolivia and Peru and another consisting of specimens from Panama. In contrast, the maximum likelihood showed different results, that specimens from Peru were more basal in the phylogenetic tree, followed by specimens from Bolivia, Ecuador and Panama (Figure 2).
Carollia perspicillata - Specimens from five different locations were analyzed (Brazilian Amazon, Minas Gerais, Brazil, Mexico, Ecuador and Suriname). In parsimony and maximum likelihood the specimens located in Brazil (Minas Gerais) are more basal, followed by two derived groups, one consisting of specimens of the Brazilian Amazon, Mexico and Suriname and another group consisting of specimens from Brazilian Amazon and Ecuador (Figure 2).
Ectophylla alba - It was analyzed specimens from two different localities (Panama and Costa Rica). In parsimony and maximum likelihood, the specimens located in Panama are basal, followed by specimens from Costa Rica. Showing dispersion towards the North to Central America.
]]> Glossophaga comissarisi - It was analyzed specimens from two countries (Mexico and Panama). In maximum parsimony and likelihood, the specimens located in Panama are basal, followed by specimens from Mexico, showing dispersion towards the North to Central America.Glossophaga soricina - Specimens from six different locations were analyzed (Panama, Paraguay, Venezuela, El Salvador, Mexico and Jamaica). In parsimony and maximum likelihood, the specimens located in Venezuela are basal, followed by specimens from Paraguay, Panama, El Salvador, Jamaica and Mexico, suggesting dispersion towards the north, from South America to North America (Figure 2).
Mesophylla macconelli - Specimens from four locations were analyzed (Ecuador, Bolivia, Colombia, French Guiana). In maximum parsimony, two sister groups assembled, one with specimens from Bolivia and Colombia with the same degree of phylogenetic relatedness and another group showing specimens from French Guiana and Ecuador, which are more derived. Regarding the analysis by maximum likelihood, the specimens from Ecuador are basal, followed by specimens from French Guiana, Bolivia and Colombia (most derived).
Platyrrhinus dorsalis - It was analyzed specimens from two different localities (Bolivia and Ecuador). In parsimony and maximum likelihood, the species have spread to Northern of South America, in which the specimens of Bolivia are more derived than the specimens from Ecuador.
Uroderma bilobatum - Specimens from three countries were analyzed (Honduras, Panama and Ecuador). In parsimony and maximum likelihood, the species have spread from Southern Central America to South America, where the specimens from Honduras are more basal, followed by specimens from Panama and Ecuador (both in the same degree of phylogenetic relatedness).
Vampyressa thyone - In maximum parsimony, the species showed dispersion to the North of Central America, from Panama to Costa Rica. However, analysis by maximum likelihood was different, North to South, from Costa Rica to Panama.
Vampyrodes caraccioli - Three specimens were analyzed in Peru, Trinidad and Tobago and Brazil. In parsimony and maximum likelihood, dispersion followed from Peru to Trinidad and Tobago and Brazil.
DISCUSSION
In the general analysis by maximum parsimony, it was noticed the most basal status of the specimen Carollia brevicauda from Peru (Cuzco) comparing specimens of C. perspicillata and C. brevicauda presented in Venezuela, proposing a possible origin of these specimens from the specimen of southern (Peru). A spread of these specimens could be possible to the north of the American Continent and eastward Brazil. It was found the same analysis result for genera Glossophaga, Ectophylla, Vampyrodes, Vampyressa and, thus, proposing a dispersion pattern to the north in several species of that family (Figure 1).
]]> This dispersion pattern can also be seen in the individual analysis, that some species of South America have a possible dispersion pattern to the north, in Central America and North America, such as the species Glossophaga soricina and Glossophaga comissarisi, which may have dispersed from South America to the North searching for better habitat and food. About Glossophaga soricina, there are two distinct strains of the species, one to the east of the Andes and the other one in Central America, North America, Jamaica and West of the Andes, which can further be classified as different species4 (Figures 2 and 3).
The Andes may also have influenced in species that have affinity for high altitudes. Kunz and Pena43 described the presence of Mesophylla macconelli in regions with altitude up to 1,032 m, Baker and Clark44 described Uroderma bilobatum to 1,800 m and Lewis and Wilson45 described Vampyressa pussila in 1,500 ft, among other species that also have some affinity with high altitudes.
The possible location and dispersion of Glossophaga may have been influenced by their ecology. For example, Glossophaga commissarisi is in a variety of habitats, such as rainforests, subtropical savannas and others. Another example is Glossophaga soricina according to their habitat and time of year, they may have a variable eating behavior, such as those in Mexico, that their feeding habits is mainly of nectar and pollen from April to June, then they change their feeding behavior to insectivorous. Alvarez et al46 has also described eating behavior of nectar and pollen during the dry season in Panama and eating fruits in the wet season.
Hoffmann and Baker5 reported the emergence of the Isthmus of Panama as one of the factors of the dispersion to the North America, which was confirmed by studies with terrestrial mammals. The authors have also reported that the same situation may have happened with the bats, they might have taken advantage of the Isthmus "bridge" for dispersal between the continents. However, the same authors have mentioned that the phylogenetic variation in Carollia may have occurred by vicariance with the rise of the Andes mountains, preventing the spread of these bats and isolating individuals on the other side of the Andes. The same idea can be inferred for bats from other groups that may have this dispersion pattern to North. In this present study, Carollia perspicillata and C. castanea showed the same pattern of dispersal to the North of the American Continent, and C. castanea seemed to have a distribution along the Andes to Panama (Figures 2A, 3A, 2B and 3B), it can be possible due to an affinity to high altitudes, as previously mentioned for other species. However, it was also noticed similarities with the dispersion pattern to the North towards Panama and South, and another pattern towards Peru and Bolivia, as shown in figure 3, which differs from the idea of isolating of Carollia by the Andes.
The same pattern of vicariance may have occurred with Uroderma bilobatum, that has presented dispersion to Ecuador in the Andes Region, with the specimens from Panama as a sister group.
Other species such as Artibeus jamaicensis have dispersed to the West, probably because of better habitat with the emergence of forest refuges during the Pleistocene, causing several species disperse to new habitats in search of food, as occurred with the genus Carollia5. A good example of vicariance is the most basal status Artibeus lituratus present in St. Vincent and the Grenadines, which may have been a case of isolation, caused for the formation of islands.
]]> As we obtained sequences of few specimens from close countries for Vampyressa thyone and Ectophylla alba, we cannot say that they have spread North to Central America, it needs better analyzes and specimens from other localities for a better theory of dispersion. However, both have a pattern in phylogenetic relatedness, where the species located in Panama are more basal than the ones found in Costa Rica, a possible spread towards the North.Regarding the species Platyrrhinus dorsalis, Velazco and Patterson8 describe that the ancestor of the genus Platyrrhinus may have arisen in the South of the Amazon River in Brazil and dispersed towards the Andes and the Amazon basin, originating several new species of the genus. According to these authors, the species were found in the Northern Andes, occupying the area of Ecuador, following the North to Venezuela. However, in our analysis we found evidence of a possible spread North to Venezuela.
The preference for specific types of habitat of various species may also influence the Phyllostomidae dispersion. Some species prefer humid and wet habitats, like the Amazon forest and other tropical forests, for exemple, Artibeus jamaicensis, Mesophylla maconelli, Platyrrhinus dorsalis and Vampyrodes caraccioli47,48,49. Others species, as example Uroderma bilobatum, prefer certain types of foods such as figs obtained in high trees of tropical forests44.
CONCLUSION
It can be concluded that the Phyllostomidade family, in general, may have a dispersion pattern on the American Continent, it can be noticed that several species, in intraspecific analysiss, have a possible pattern, in the North America, as Carollia brevicauda, C. castanea, C. perspicillata, Glossophaga comissarisi and G. soricina or to the South, as Uroderma bilobatum and Vampyrodes caraccioli. Regarding the other species analyzed, it cannot be suggested a dispersion pattern, because the analisis presented a random dispersion or few specimens from different and distant localities.
Many other studies work at genus and species level, rarely at family level, due to lack of specimens from distant and different localities. Here it was performed an analysis of GenBank sequences for a possible dispersion theory to the North of the American Continent for Phyllostomidae family. The reduced number of retrieved sequences were low because of the lack of published sequences with its exact location of the specimen. However, the sequences obtained were really important due to the distinct and distant locations and features that can prove other dispersion theories, as the vicariance occurred by the emergence of the Andes Mountains. Thus, it concludes that the family may have a dispersion pattern (probably to the North), however other analysis must be performed in the future, as phylogenetic regression, molecular clock and the discovery of new sequences at other localities in the American Continent.
ACKNOWLEDGENTS
To Clayton Lima and the Centro de Inovações Tecnológicas, Arbovirologia e Febres Hemorrágicas and Meio Ambiente sections from Instituto Evandro Chagas and the Universidade Federal do Pará.
]]>REFERENCES
1 Freygang CC. Estudos filogenéticos dos morcegos filostomídeos da Região Neotropical [tese]. Porto Alegre (RS): Universidade Federal do Rio Grande do Sul; 2006.
2 Reis NR, Peracchi AL, Pedro WA, Lima IP Morcegos do Brasil. Londrina: Universidade Estadual de Londrina; 2007. 253 p.
3 Hoffmann Jauge FG. Systematics and phylogeography of neotropical bats of the genera Carollia, Glossophaga, Tonatia and Uroderma [thesis]. Lubbock (TX): Texas Tech University; 2002. 244 p. [Link]
4 Hoffmann FG, Baker RJ. Systematics of bats of the genus Glossophaga (Chiroptera: Phyllostomidae) and phylogeography in G. soricina based on the cytochrome-b gene. J Mammal. 2001 Nov;82(4):1092-101. Doi: 10.1644/1545-1542(2001)082<1092:SOBOTG>2.0.CO;2 [Link]
5 Hoffmann FG, Baker RJ. Comparative phylogeography of short-tailed bats (Carollia: Phyllostomidae). Mol Ecol. 2003 Dec;12(12):3403-14. Doi: 10.1046/j.1365-294X.2003.02009.x [Link]
6 Larsen PA, Hoofer SR, Bozeman MC, Pedersen SC, Genoways HH, Phillips CJ, et al. Phylogenetics and phylogeography of the Artibeus jamaicensis complex bases on cytochrome-b DNA sequences. J Mammal. 2007 Jun;88(3):712-7. Doi: 10.1644/06-MAMM-A-125R.1 [Link]
7 Martins FM, Templeton AR, Pavan AC, Kohlbach BC, Morgante JS. Phylogeography of the common vampire bat (Desmodusrotundus): marked population structure, Neotropical Pleistocene vicariance and incongruence between nuclear and mtDNA markers. BMC Evol Biol. 2009 Dec;9(294):1-13. Doi: 10.1186/1471-2148-9-294 [Link]
8 Velazco PM, Patterson BD. Phylogenetics and biogeography of the broad-nosed bats genus Platyrrhinus (Chiroptera: Phyllostomidae). Mol Phylogenet Evol. 2008 Dec;49(3):749-59. Doi: 10.1016/j.ympev.2008.09.015 [Link]
]]> 9 Simmons NB. Order Chiroptera. In: Wilson DE, Reeder DM. Mammal species of the world: a taxonomic and geographic reference. 3th ed. Vol. 1. Baltimore: Johns Hopkins University Press; 2005. p. 312-529.10 Springer MS, Teeling EC, Madsen O, Stanhope MJ, Jong WW. Integrated fossil and molecular data reconstruct bat echolocation. Proc Natl Acad Sci USA. 2001 May;98(11):6241-6. Doi: 10.1073/pnas.111551998 [Link]
11 Teeling EC, Madsen O, Van den Bussche RA, Jong WW, Stanhope MJ, Springer MS. Microbatparaphyly and the convergent evolution ofa key innovation in Old World rhinolophoid microbats. Proc Natl Acad Sci USA. 2002 Feb;99(3):1431-6. Doi: 10.1073/pnas.022477199 [Link]
12 Teeling EC, Springer MS, Madsen O, Bates P, O'Brien SJ, Murphy WJ. A molecular phylogeny for bats illuminates biogeography and the fossil record. Science. 2005 Jan;307(5709):580-4. Doi: 10.1126/science.1105113 [Link]
13 Nowak RM. Walkers mammals of the world. 6th ed. Baltimore: The Johns Hopkins University Press; 1999. 1936 p.
14 Baker RJ, Porter CA, Patton JC, Bussche RAV. Systematics of bats of the family phyllostomidae based on Rag2 DNA sequences. Occas Pap Tex Tech Univ Mus. 2000;202:1-16. [Link]
15 Ribeiro NAB. Evolução cromossômica e filogenia de quatro espécies de morcegos nectarívoros do Novo Mundo (Phyllostomidae-Chiroptera) [dissertação]. Belém (PA): Universidade Federal do Pará, Centro de Ciências Biológicas; 2000.
16 Wetterer AL, Rockman MV, Simmons NB. Phylogeny of phyllostomid bats (Mammalia: Chiroptera): data from diverse morphological systems, sex chromosomes, and restriction sites. Bull Am Mus Nat Hist. 2000 Mar;248:1-200. [Link]
17 Jones KE, Purvis A, Maclarnon A, Bininda-Emonds ORP, Simmons NB. A phylogenetic supertree of the bats (Mammalia: Chiroptera). Biol Rev Camb Philos Soc. 2002 May;77(2):223-59. Doi: 10.1017/S1464793101005899 [Link]
18 Baker RJ. Biology of bats of the New World family Phyllostomatidae. Spec Publ Mus Tex Tech Univ. 1979;16:1-441.
]]> 19 Koopman KF. Order chiroptera. In: Wilson DE, Reeder DM. Mammal species of the world, a taxonomic and geographic reference. 2nd ed. Washington: Smithsonian Institute Press; 1993. p. 137-241.20 Mckenna MC, Bell SK. Classification of mammals above the species level. New York: Columbia University Press; 1997. 631 p.
21 Baker RJ, Hoofer SR, Porter CA, Van den Bussche RA. Diversification among New World leaf-nosed bats: an evolutionary hypothesis and classification inferred from digenomic congruence of DNA sequence. Occas Pap Tex Tech Univ Mus. 2003 Dec;230:1-32. [Link]
22 Baker RJ. Comparative cytogenetics of the New World leaf-nosed bats (Phyllostomidae). Period Biol. 1973;75:37-45. [Link]
23 Brown WM, George MJR, Wilson AC. Rapid evolution of animal mitochondrial DNA. Proc Natl Acad Sci USA. 1979 Apr;76(4):1967-71. [Link]
24 Snustad DP, Simmons MJ. Fundamentos de genética. 4. ed. Rio de Janeiro: Guanabara Koogan; 2008. Genética de organelas e cloroplastos; p. 579-86.
25 Irwin DM, Kocher TD, Wilson AC. Evolution of the cytochrome b gene in mammals. J Mol Evol. 1991 Feb;32(2):128-44. [Link]
26 Farias IP, Ortí G, Sampaio I, Schneider H, Meyer A. The cytochrome b gene as a phylogenetic marker: the limits of resolution for analyzing relationship among cichlid fishes. J Mol Evol. 2001 Aug;53(2):89-103. Doi: 10.1007/s002390010197 [Link]
27 Beheregaray LB. Twenty years of phylogeography: the state of the field and the challenges for the Southern Hemisphere. Mol Ecol. 2008 Sep;17(17):3754-74. Doi: 10.1111/j.1365-294X.2008.03857.x [Link]
28 Popa-Lisseanu AG, Voigt CC. Bats on the move. J Mammal. 2009 Dec;90(6):1283-9. Doi: 10.1644/09-MAMM-S-130R2.1 [Link]
]]> 29 Esbérard CEL, Lima IP, Nobre PH, Althoff SL, Jordão-Nogueira T, Dias D, et al. Evidence of vertical migration in the Ipanema bat Pygoderma bilabiatum (Chiroptera: Phyllostomidae: Stenodermatinae). Zoologia. 2011 Dec;28(6):717-24. Doi: 10.1590/S1984-46702011000600004 [Link]30 Hedenström A. Optimal migration strategies in bats. J Mammal. 2009 Dec;90(6):1298-309. Doi: 10.1644/09-MAMM-S-075R2.1 [Link]
31 Mcguire LP. Physiological ecology of bat migration [thesis]. London (ON): University of Western Ontario; 2012.
32 Montalvo LGH. Evidence of altitudinal movements of Leptonycteris curasoae (Chiroptera: Phyllostomidae) in central Mexico. Rev Mexicana Mastozool. 1997 ene;2(1):116-8. [Link]
33 Van den Bussche RA, Hudgeons JL, Baker RJ. Phylogenetic accuracy, stability, and congruence: relationships within and among the New World bat genera Artibeus, Dermanura and Koop-mania. In: Kunz TH, Racey PA, editors. Bat biology and conservation. Washington: Smithsonian Institution Press; 1998. p. 59-71.
34 Hoofer SR, Solari S, Larsen PA, Bradley RD, Baker RJ. Phylogenetics of the fruit-eating bats (Phyllostomidae: Artibeina) inferred from mitochondrial DNA sequences. Occas Pap Tex Tech Univ Mus. 2008 Oct;176:1-15. [Link]
35 Larsen PA, Marchan-Rivadeneira R, Baker RJ. Speciation dynamics of the fruit-eating bats (genus artibeus): with evidence of ecological divergence in Central American populations. In: Adams RA, Pedersen SC, editors. Bat evolution, ecology, and conservation. New York: Springer; 2013. p. 315-39.Doi: 10.1007/978-1-4614-7397-8_16 [Link]
36 Wright AJ, Van den Bussche RA, Burton K, Lim M, Engstrom D, Baker RJ. Systematics of the genera Carollia and Rhinophylla based on the cytochrome-B gene. J Mammal. 1999 Nov;80(4):1202-13.[Link]
37 Baker RJ, Solari S, Hoffmann FG. A new Central American species from the Carollia brevicauda complex. Occas Pap Tex Tech Univ Mus. 2002 Oct;217:1-12. [Link]
38 Pavan AC, Martins F, Santos FR, Ditchfield A, Redondo RAF. Patterns of diversification in two species of short-tailed bats (Carollia Gray, 1838): the effects of historical fragmentation of Brazilian rainforests. Biol J Linnean Soc. 2011 Mar;102(3):527-39. Doi: 10.1111/j.1095-8312.2010.01601.x [Link]
]]> 39 Hoofer SR, Baker RJ. Molecular systematics of Vampyressine bats (Phyllostomidae: Stenodermatinae) with comparison of direct and indirect surveys of mitochondrial DNA variation. Mol Phylogenet Evol. 2006 May;39(2):424-38. Doi: 10.1016/j.ympev.2005.12.001 [Link]40 Porter CA, Baker RJ. Systematics of Vampyressa and related genera of Phyllostomid bats as determined by cytochrome-b sequences. J Mammal. 2004 Feb;85(1):126-32.Doi: 10.1644/BWG-110 [Link]
41 Hoffmann FG, Owen JG, Baker RJ. mtDNA perspective of chromosomal diversification and hybridization in Peters' tent-making bat (Uroderma bilobatum: Phyllostomidae). Mol Ecol. 2003 Nov;12(11):2981-93. Doi: 10.1046/j.1365-294X.2003.01959.x [Link]
42 Velazco PM, Simmons NB. Systematics and taxonomy of great striped-faced bats of the genus Vampyrodes Thomas, 1990 (Chiroptera: Phyllostomidae). Am Mus Novit. 2011 Apr;3710:1-35. [Link]
43 Kunz TH, Pena IM. Mesophylla macconnelli. Mamm Species.1992 Dec;405:1-5.[Link]
44 Baker RJ, Clark CL. Uroderma bilobatum. Mamm Species. 1987 Feb;279:1-4.[Link]
45 Lewis SE, Wilson DE. Vampyress apusilla. Mamm Species. 1987 Aug;292:1-5. [Link]
46 Alvarez J, Willig MR, Jones JK, Webster D. Glossophaga soricina. Mamm Species. 1991 Nov;379:1-7.[Link]
47 Willis KB, Willig MR, Jones JK. Vampyrodes caraccioli. Mamm Species. 1990 Oct;359:1-4. [Link]
48 Kunz TH, Pena IM. Mesophylla macconnelli. Mamm Species.1992 Dec;405:1-5. [Link]
]]> 49 Ortega J, Castro-Arellano I. Artibeus jamaicensis. Mamm Species. 2001 Jun;662:1-9. [Link]
 Correspondence / Correspondência / Correspondencia:
Correspondence / Correspondência / Correspondencia:
Gustavo Moraes Holanda
Rua Antonio Barreto, 747. Bairro: Umarizal
CEP: 66055-050 Belém-Pará-Brasil
Phone #: +55 (91) 8757-2970
E-mail: holandagm@gmail.com
Received / Recebido em / Recibido en: 10/1/2012 ]]> Accepted / Aceito em / Aceito en: 19/6/2012
]]>